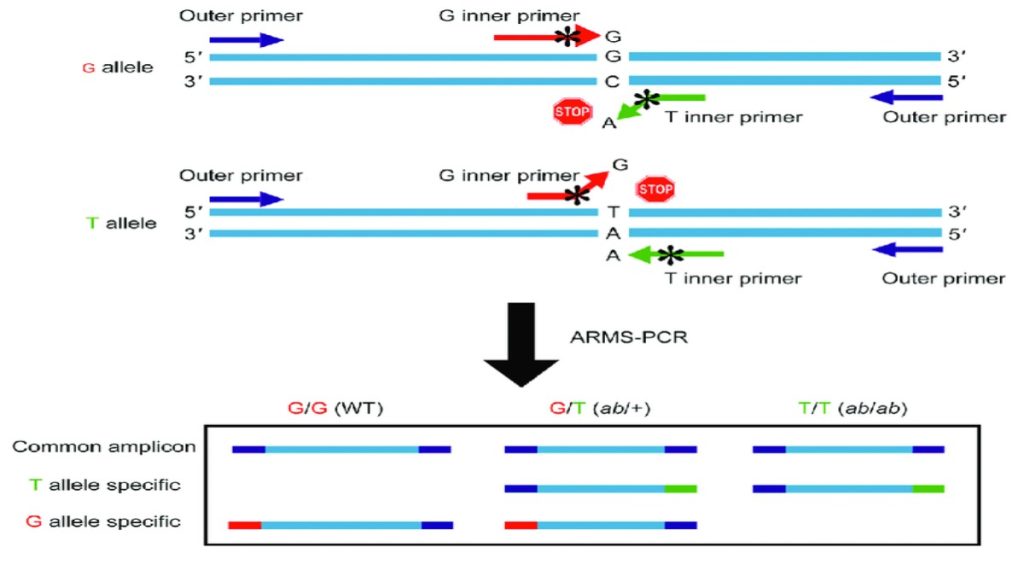
The allele-specific PCR also called as an ARMS- PCR (amplification refractory mutation system) or PASA (PCR amplification of specific alleles) or AS-PCR used to detect the SNPs.
More specifically, it is adopted to detect the known SNPs (single nucleotide polymorphism), however, we cannot identify new mutations by ARMS-PCR.
Kary Mullis described the technique of in vitro amplification in the year 1983. After a few years of the discovery of the actual PCR technique, C. R. Newton and coworkers discovered the ARMS-PCR or allele-specific PCR technique.
The technique is majorly used for the genotyping of the single nucleotide polymorphism with the help of the refractory primers.
In this article, we will understand the allele-specific PCR/ ARMS-PCR and its importance in medical science.
Further, we will explain the mechanism of how we can develop different primers for allele-specific PCR.
Before going to the actual process lets first understand the terms “Allele-specific PCR” and “ARMS-PCR.”
What is allele-specific PCR?
The term suggests that the technique used in this type of PCR is specific to the particular type of allele. An allele is the alternative form of a gene.
If one allele has an SNP and the other alternative form is normal, we can analyse both the alleles by designing specific primers for each allele.
For that, we have to modify the single base at the 3’ end of the primer (one primer matches the normal allele and one primer matches the mutant allele).
The PCR is performed simultaneously in a single reaction. If a mutant allele is present, then the PCR amplifies the mutant allele or if the normal allele is present, the normal allele will amplify.
What is ARMS-PCR?
The allele-specific PCR is also called as the (amplification refractory mutation system) ARMS-PCR because of the use of two different primers for two different alleles.
Here the word “refractory” is very important (Refractory= resistant to something).
As we discussed, two sets of primers are designed, the mutant set of the primer is refractory (resistant) to the normal PCR and the normal set of the primers are refractory to the mutant PCR reaction.
That is why it is called an amplification refractory mutation system. The name ARMS-PCR is given by its actual developer C. R. Newton.
The principle of allele-specific ARMS-PCR:
The mechanism of the ARMS PCR is based on the modification of the primers for different alleles.
Here, the 3’ end of the primers is modified in such a way that one set of the primer can amplify the normal allele and others can amplify the mutant allele. The mismatch single base is introduced at the 3’ end of the primer. This mismatch allows the primer to amplify one single allele.
The concept of mismatch:
Here the mismatch between the primer and the template DNA plays a crucial role in achieving the amplification.
Introduction of a mismatch at the 3’ end of the primer alters the annealing temperature for that particular allele.
Strong mismatch: G/A, C/T, T/T
Medium mismatch: A/A, G/G, C/C,
Weak mismatch: C/A, G/T

Due to the absence of the exonuclease activity of Taq DNA polymerase, the mismatch cannot be repaired. Why high fidelity Taq DNA polymerase does not use in the ARMS PCR? we will answer this question in the last section of this article.