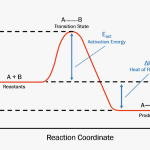
A substance that helps a chemical reaction to occur is called a catalyst, and the molecules that catalyze biochemical reactions are called enzymes. Most enzymes are proteins and perform the critical task of lowering the activation energies of chemical reactions inside the cell. Most of the reactions critical to a living cell happen too slowly at normal temperatures to be of any use to the cell. Without enzymes to speed up these reactions, life could not persist. Enzymes do this by binding to the reactant molecules and holding them in such a way as to make the chemical bond-breaking and -forming processes take place more easily. It is important to remember that enzymes do not change whether a reaction is exergonic (spontaneous) or endergonic. This is because they do not change the free energy of the reactants or products. They only reduce the activation energy required for the reaction to go forward (Figure 4.7). In addition, an enzyme itself is unchanged by the reaction it catalyzes. Once one reaction has been catalyzed, the enzyme is able to participate in other reactions.
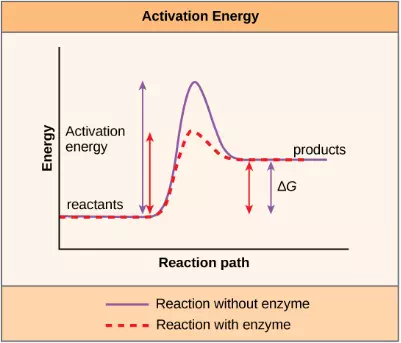
Figure 4.7 Enzymes lower the activation energy of the reaction but do not change the free energy of the reaction.
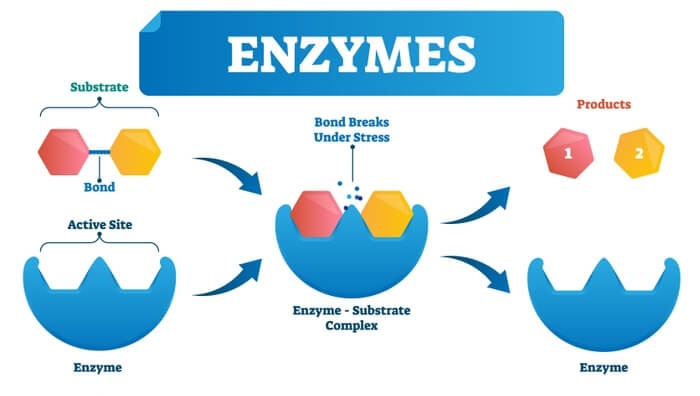
The chemical reactants to which an enzyme binds are called the enzyme’s substrates. There may be one or more substrates, depending on the particular chemical reaction. In some reactions, a single reactant substrate is broken down into multiple products. In others, two substrates may come together to create one larger molecule. Two reactants might also enter a reaction and both become modified, but they leave the reaction as two products. The location within the enzyme where the substrate binds is called the enzyme’s active site. The active site is where the “action” happens. Since enzymes are proteins, there is a unique combination of amino acid side chains within the active site. Each side chain is characterized by different properties. They can be large or small, weakly acidic or basic, hydrophilic or hydrophobic, positively or negatively charged, or neutral. The unique combination of side chains creates a very specific chemical environment within the active site. This specific environment is suited to bind to one specific chemical substrate (or substrates).
Active sites are subject to influences of the local environment. Increasing the environmental temperature generally increases reaction rates, enzyme-catalyzed or otherwise. However, temperatures outside of an optimal range reduce the rate at which an enzyme catalyzes a reaction. Hot temperatures will eventually cause enzymes to denature, an irreversible change in the three-dimensional shape and therefore the function of the enzyme. Enzymes are also suited to function best within a certain pH and salt concentration range, and, as with temperature, extreme pH, and salt concentrations can cause enzymes to denature.
For many years, scientists thought that enzyme-substrate binding took place in a simple “lock and key” fashion. This model asserted that the enzyme and substrate fit together perfectly in one instantaneous step. However, current research supports a model called induced fit (Figure 4.8). The induced-fit model expands on the lock-and-key model by describing a more dynamic binding between enzyme and substrate. As the enzyme and substrate come together, their interaction causes a mild shift in the enzyme’s structure that forms an ideal binding arrangement between enzyme and substrate.
When an enzyme binds its substrate, an enzyme-substrate complex is formed. This complex lowers the activation energy of the reaction and promotes its rapid progression in one of multiple possible ways. On a basic level, enzymes promote chemical reactions that involve more than one substrate by bringing the substrates together in an optimal orientation for reaction. Another way in which enzymes promote the reaction of their substrates is by creating an optimal environment within the active site for the reaction to occur. The chemical properties that emerge from the particular arrangement of amino acid R groups within an active site create the perfect environment for an enzyme’s specific substrates to react.
The enzyme-substrate complex can also lower activation energy by compromising the bond structure so that it is easier to break. Finally, enzymes can also lower activation energies by taking part in the chemical reaction itself. In these cases, it is important to remember that the enzyme will always return to its original state by the completion of the reaction. One of the hallmark properties of enzymes is that they remain ultimately unchanged by the reactions they catalyze. After an enzyme has catalyzed a reaction, it releases its product(s) and can catalyze a new reaction.
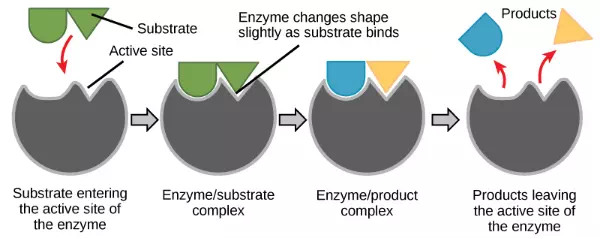
Figure 4.8 The induced-fit model is an adjustment to the lock-and-key model and explains how enzymes and substrates undergo dynamic modifications during the transition state to increase the affinity of the substrate for the active site.
It would seem ideal to have a scenario in which all of an organism’s enzymes existed in abundant supply and functioned optimally under all cellular conditions, in all cells, at all times. However, a variety of mechanisms ensures that this does not happen. Cellular needs and conditions constantly vary from cell to cell, and change within individual cells over time. The required enzymes of stomach cells differ from those of fat storage cells, skin cells, blood cells, and nerve cells. Furthermore, a digestive organ cell works much harder to process and break down nutrients during the time that closely follows a meal compared with many hours after a meal. As these cellular demands and conditions vary, so must the amounts and functionality of different enzymes.
Since the rates of biochemical reactions are controlled by activation energy, and enzymes lower and determine activation energies for chemical reactions, the relative amounts and functioning of the variety of enzymes within a cell ultimately determine which reactions will proceed and at what rates. This determination is tightly controlled in cells. In certain cellular environments, enzyme activity is partly controlled by environmental factors like pH, temperature, salt concentration, and, in some cases, cofactors or coenzymes.
Enzymes can also be regulated in ways that either promote or reduce enzyme activity. There are many kinds of molecules that inhibit or promote enzyme function, and various mechanisms by which they do so. In some cases of enzyme inhibition, an inhibitor molecule is similar enough to a substrate that it can bind to the active site and simply block the substrate from binding. When this happens, the enzyme is inhibited through competitive inhibition, because an inhibitor molecule competes with the substrate for binding to the active site.
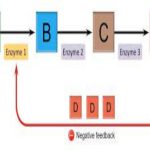
On the other hand, in noncompetitive inhibition, an inhibitor molecule binds to the enzyme in a location other than the active site, called an allosteric site, but still manages to block substrate binding to the active site. Some inhibitor molecules bind to enzymes in a location where their binding induces a conformational change that reduces the affinity of the enzyme for its substrate. This type of inhibition is called allosteric inhibition (Figure 4.9). Most allosterically regulated enzymes are made up of more than one polypeptide, meaning that they have more than one protein subunit. When an allosteric inhibitor binds to a region on an enzyme, all active sites on the protein subunits are changed slightly such that they bind their substrates with less efficiency. There are allosteric activators as well as inhibitors. Allosteric activators bind to locations on an enzyme away from the active site, inducing a conformational change that increases the affinity of the enzyme’s active site(s) for its substrate(s) (Figure 4.9).
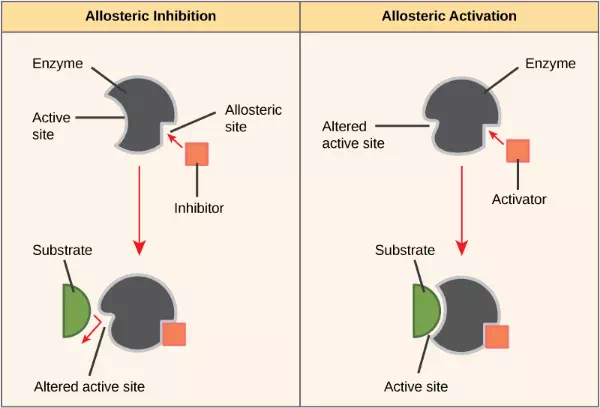
Figure 4.9 Allosteric inhibition works by indirectly inducing a conformational change to the active site such that the substrate no longer fits. In contrast, in allosteric activation, the activator molecule modifies the shape of the active site to allow a better fit of the substrate.
Pharmaceutical Drug Developer

Figure 4.10 Have you ever wondered how pharmaceutical drugs are developed?
Enzymes are key components of metabolic pathways. Understanding how enzymes work and how they can be regulated are key principles behind the development of many of the pharmaceutical drugs on the market today. Biologists working in this field collaborate with other scientists to design drugs (Figure 4.10).
Consider statins for example—statins is the name given to one class of drugs that can reduce cholesterol levels. These compounds are inhibitors of the enzyme HMG-CoA reductase, which is the enzyme that synthesizes cholesterol from lipids in the body. By inhibiting this enzyme, the level of cholesterol synthesized in the body can be reduced. Similarly, acetaminophen, popularly marketed under the brand name Tylenol, is an inhibitor of the enzyme cyclooxygenase. While it is used to provide relief from fever and inflammation (pain), its mechanism of action is still not completely understood.
How are drugs discovered?
One of the biggest challenges in drug discovery is identifying a drug target. A drug target is a molecule that is literally the target of the drug. In the case of statins, HMG-CoA reductase is the drug target. Drug targets are identified through painstaking research in the laboratory. Identifying the target alone is not enough; scientists also need to know how the target acts inside the cell and which reactions go awry in the case of disease. Once the target and the pathway are identified, then the actual process of drug design begins. In this stage, chemists and biologists work together to design and synthesize molecules that can block or activate a particular reaction. However, this is only the beginning: If and when a drug prototype is successful in performing its function, then it is subjected to many tests from in vitro experiments to clinical trials before it can get approval from the U.S. Food and Drug Administration to be on the market.
Many enzymes do not work optimally, or even at all, unless bound to other specific non-protein helper molecules. They may bond either temporarily through ionic or hydrogen bonds, or permanently through stronger covalent bonds. Binding to these molecules promotes optimal shape and function of their respective enzymes. Two examples of these types of helper molecules are cofactors and coenzymes. Cofactors are inorganic ions such as ions of iron and magnesium. Coenzymes are organic helper molecules, those with a basic atomic structure made up of carbon and hydrogen. Like enzymes, these molecules participate in reactions without being changed themselves and are ultimately recycled and reused. Vitamins are the source of coenzymes. Some vitamins are the precursors of coenzymes and others act directly as coenzymes. Vitamin C is a direct coenzyme for multiple enzymes that take part in building the important connective tissue, collagen. Therefore, enzyme function is, in part, regulated by the abundance of various cofactors and coenzymes, which may be supplied by an organism’s diet or, in some cases, produced by the organism.
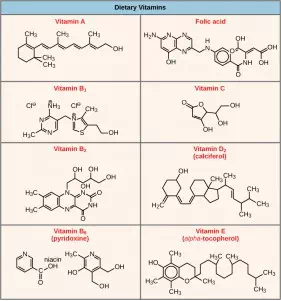
Figure 4.11 Vitamins are important coenzymes or precursors of coenzymes, and are required for enzymes to function properly. Multivitamin capsules usually contain mixtures of all the vitamins at different percentages.